Frontiers | Proteomics Approaches for Biomarker and Drug Target Discovery in ALS and FTD | Neuroscience

PDF) Data-Independent Acquisition Mass Spectrometry-Based Proteomics and Software Tools: A Glimpse in 2020
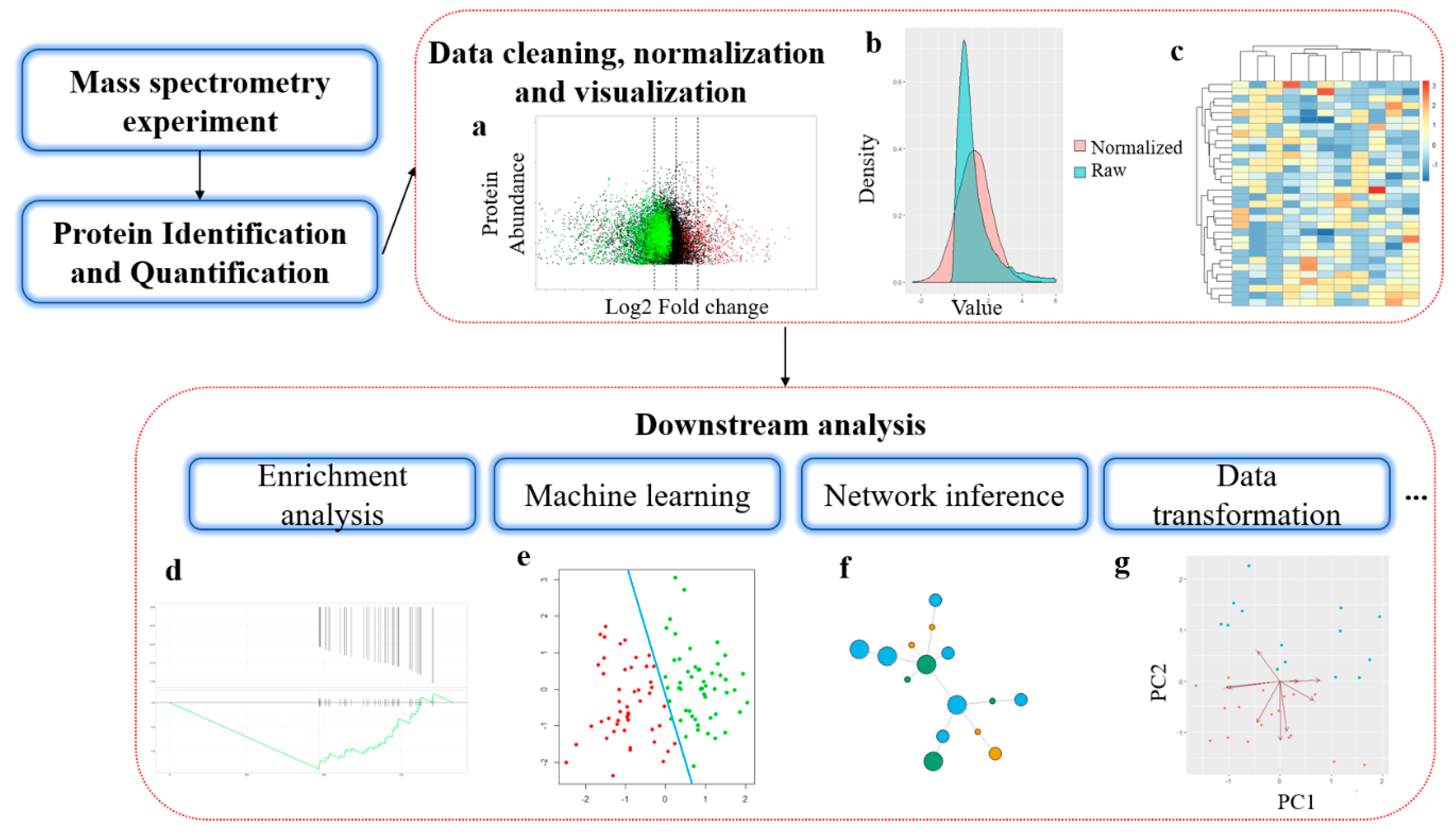
IJMS | Free Full-Text | Bioinformatics Methods for Mass Spectrometry-Based Proteomics Data Analysis | HTML
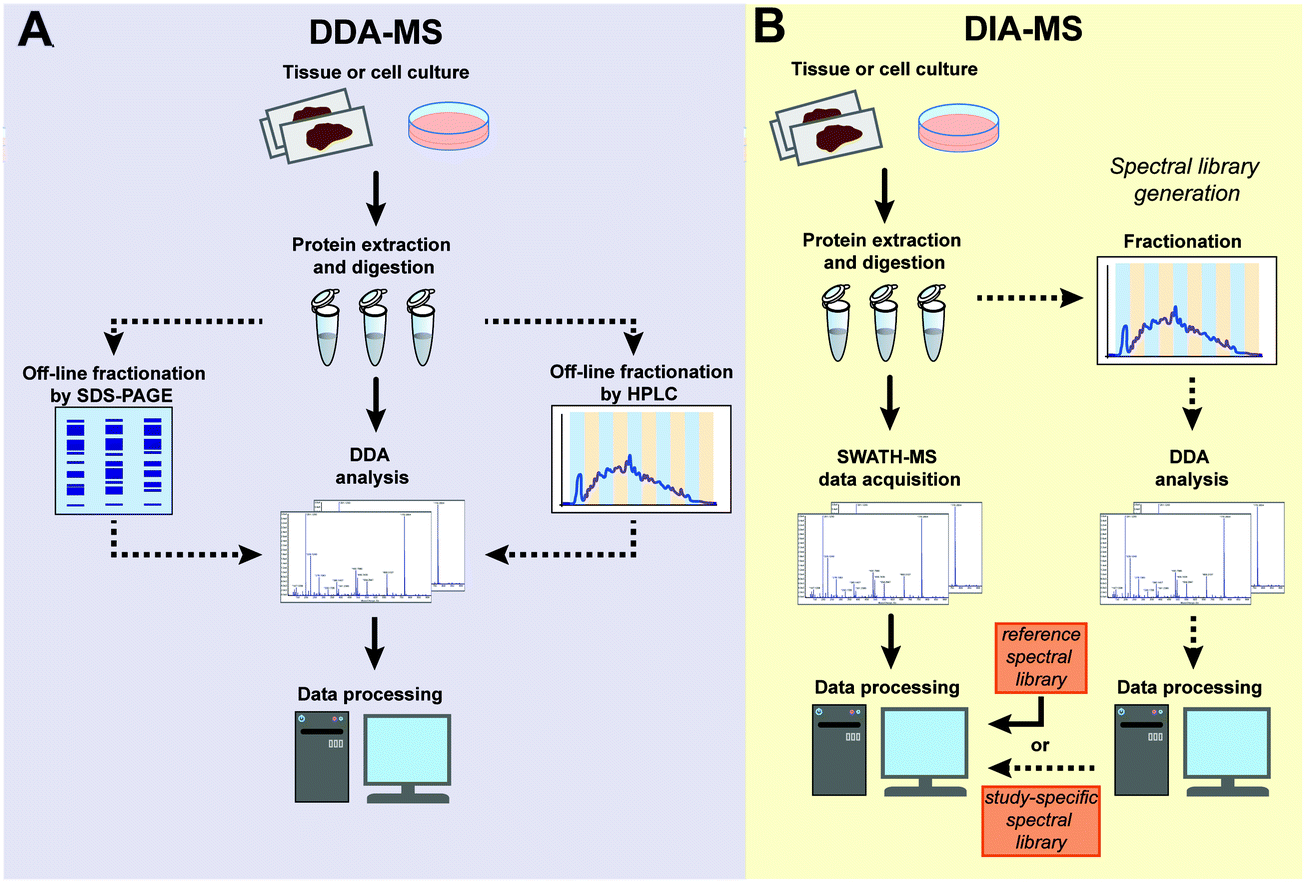
Data-independent acquisition mass spectrometry (DIA-MS) for proteomic applications in oncology - Molecular Omics (RSC Publishing) DOI:10.1039/D0MO00072H
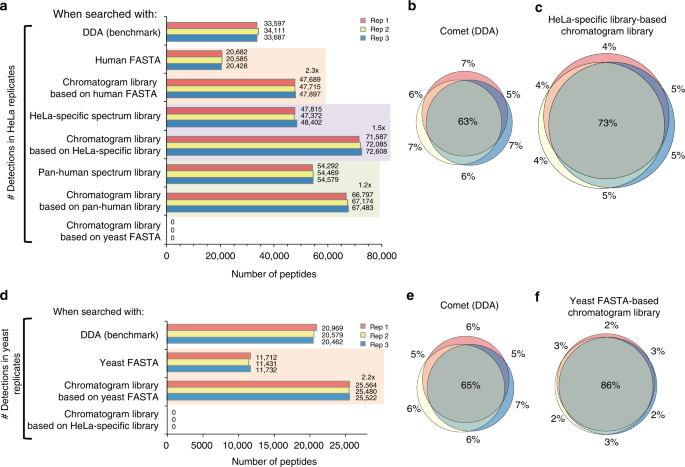
Chromatogram libraries improve peptide detection and quantification by data independent acquisition mass spectrometry | Nature Communications
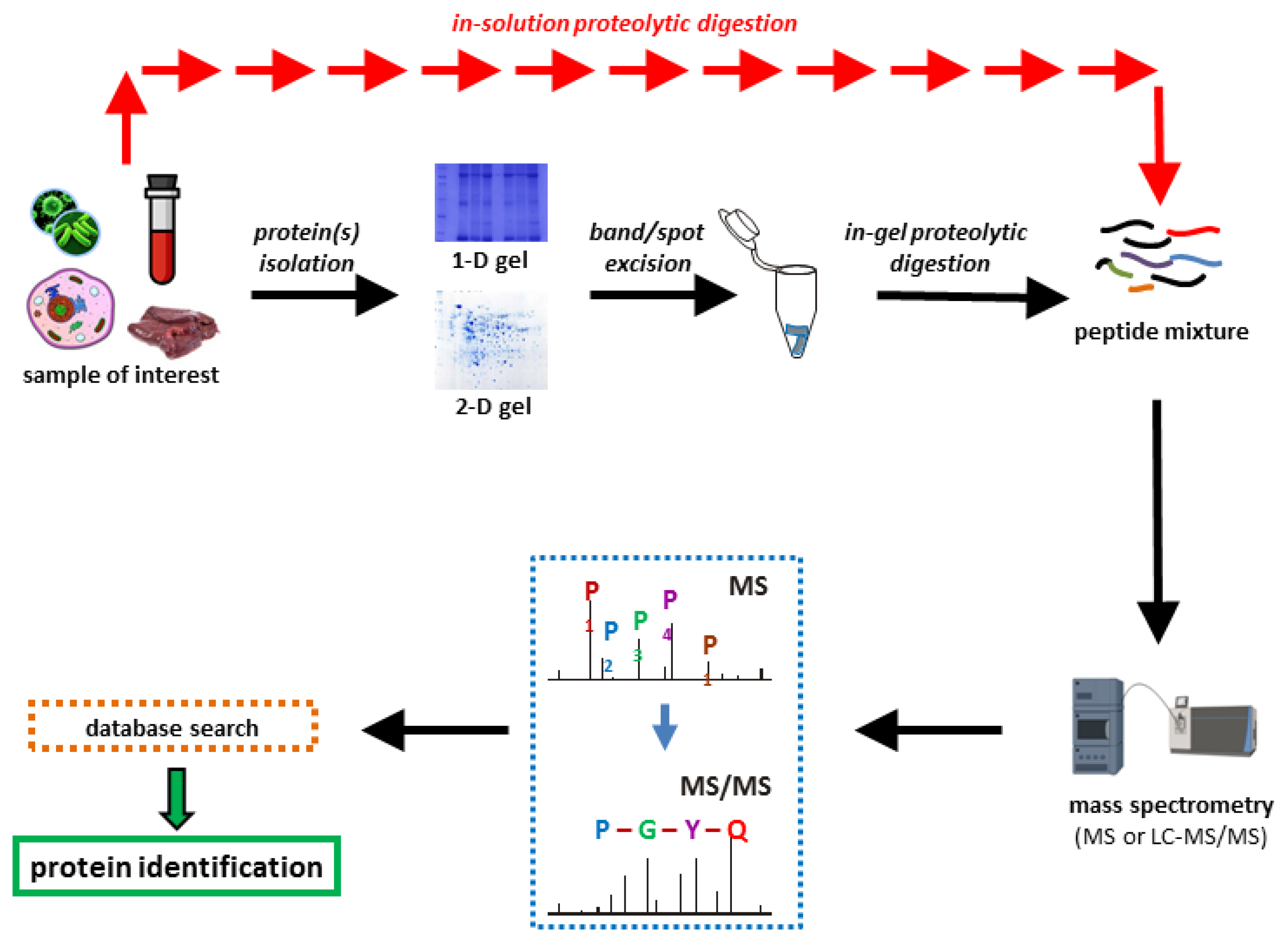
Proteomes | Free Full-Text | A Critical Review of Bottom-Up Proteomics: The Good, the Bad, and the Future of This Field | HTML

DDIA: data dependent-independent acquisition proteomics - DDA and DIA in a single LC-MS/MS run | bioRxiv

New Nature Communications publication by Mann & Theis Groups harnesses the benefits of large-scale peptide collisional cross section (CCS) measurements and deep learning for 4D-proteomics

Data‐independent acquisition‐based SWATH‐MS for quantitative proteomics: a tutorial | Molecular Systems Biology

Hybrid Spectral Library Combining DIA-MS Data and a Targeted Virtual Library Substantially Deepens the Proteome Coverage - ScienceDirect
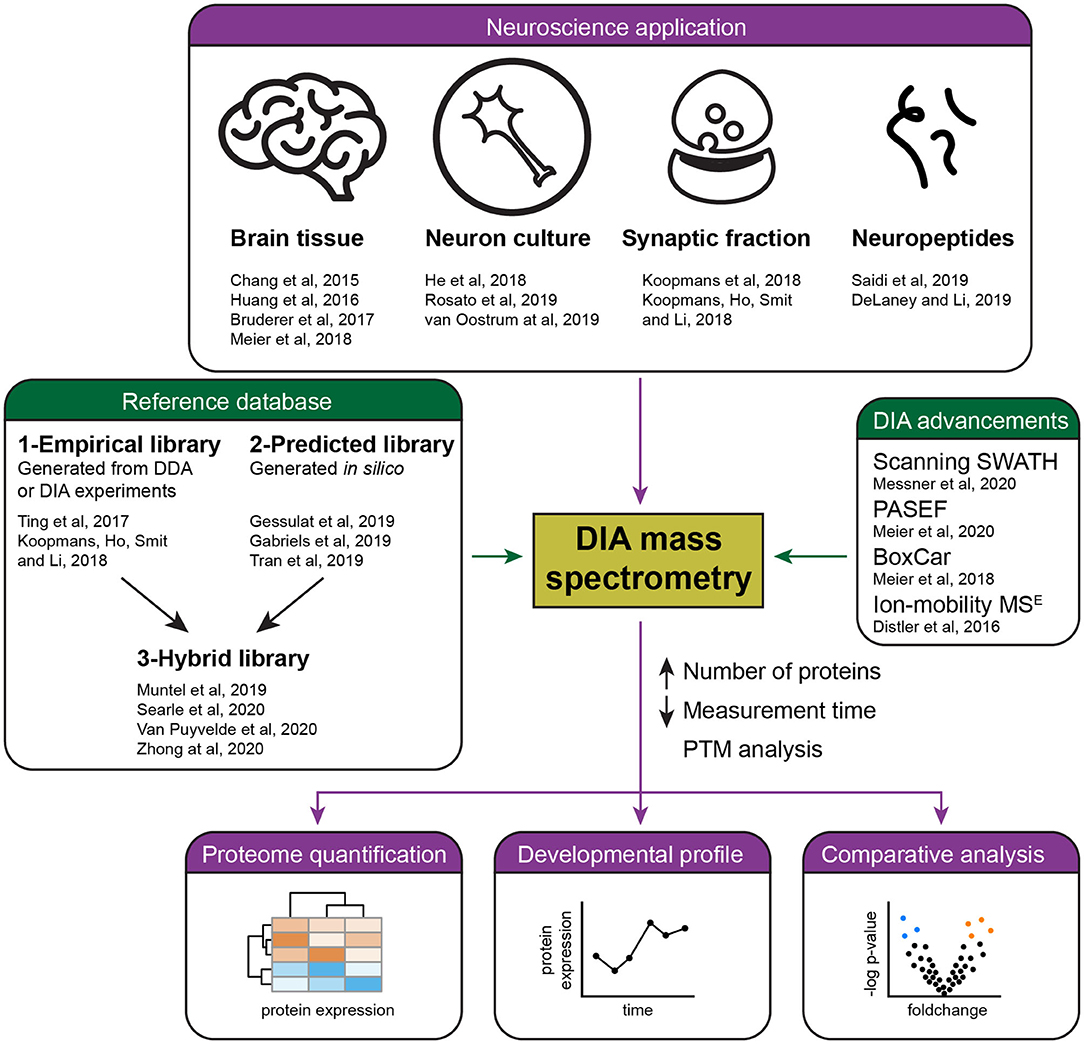
Frontiers | Recent Developments in Data Independent Acquisition (DIA) Mass Spectrometry: Application of Quantitative Analysis of the Brain Proteome | Molecular Neuroscience
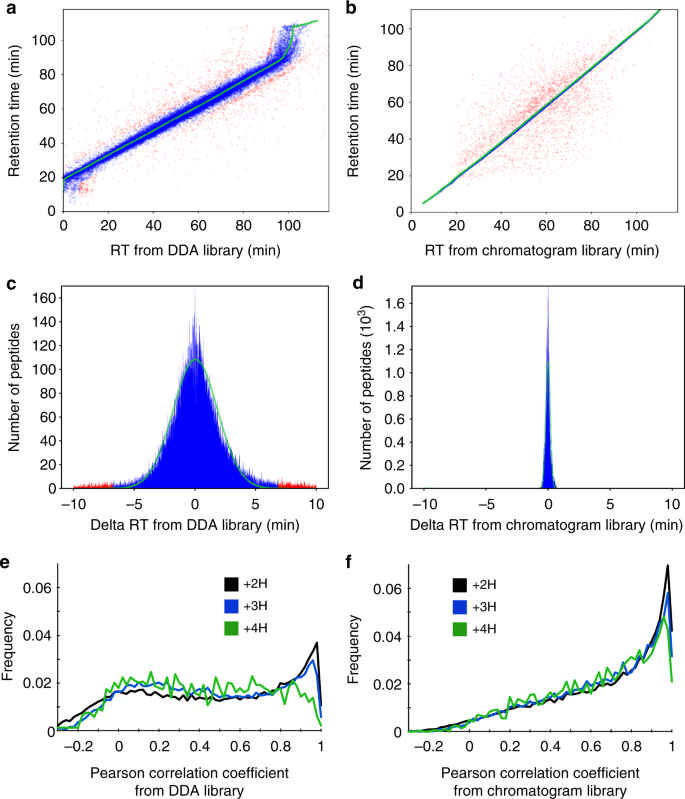
Chromatogram libraries improve peptide detection and quantification by data independent acquisition mass spectrometry | Nature Communications

Sensitive Immunopeptidomics by Leveraging Available Large-Scale Multi-HLA Spectral Libraries, Data-Independent Acquisition, and MS/MS Prediction - Molecular & Cellular Proteomics

Group-DIA: analyzing multiple data-independent acquisition mass spectrometry data files | Nature Methods

Data‐Independent Acquisition Mass Spectrometry‐Based Proteomics and Software Tools: A Glimpse in 2020 - Zhang - 2020 - PROTEOMICS - Wiley Online Library
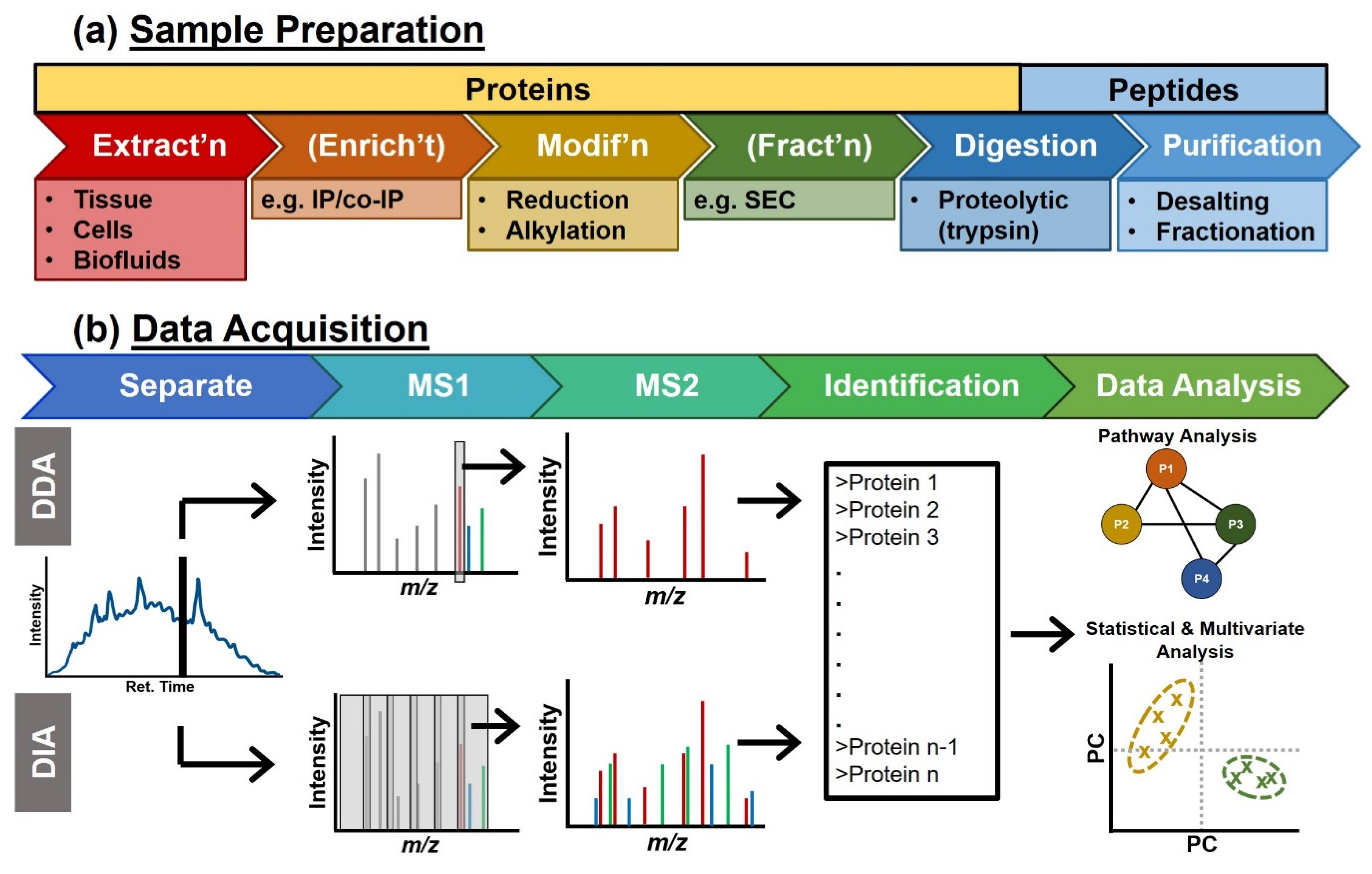
Molecules | Free Full-Text | Mass Spectrometry Advances and Perspectives for the Characterization of Emerging Adoptive Cell Therapies | HTML

Data-independent acquisition schemes in bottom-up proteomics. (1) LC... | Download Scientific Diagram

DIA-NN: neural networks and interference correction enable deep proteome coverage in high throughput | Nature Methods
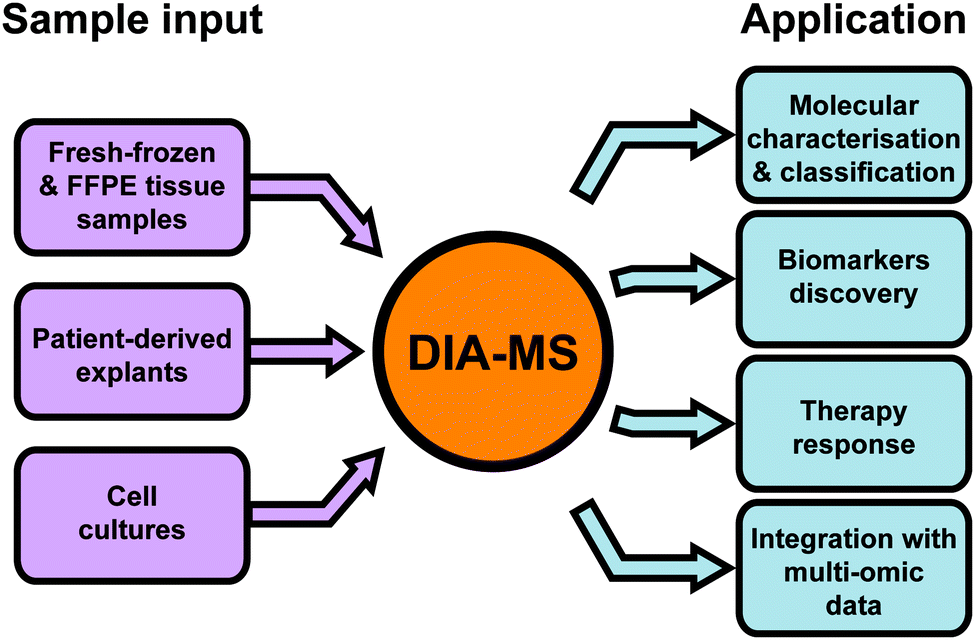
Data-independent acquisition mass spectrometry (DIA-MS) for proteomic applications in oncology - Molecular Omics (RSC Publishing) DOI:10.1039/D0MO00072H
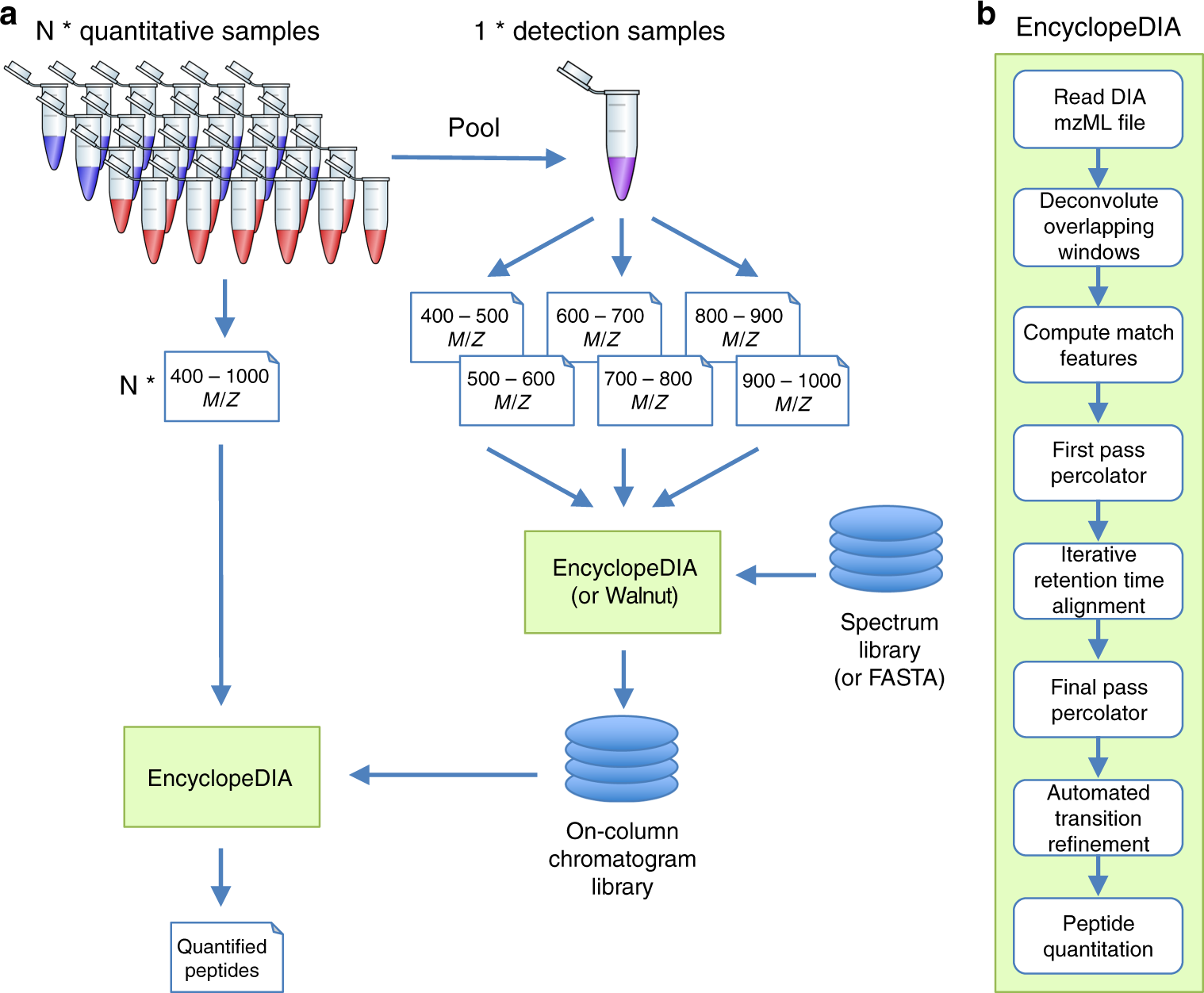
Chromatogram libraries improve peptide detection and quantification by data independent acquisition mass spectrometry | Nature Communications